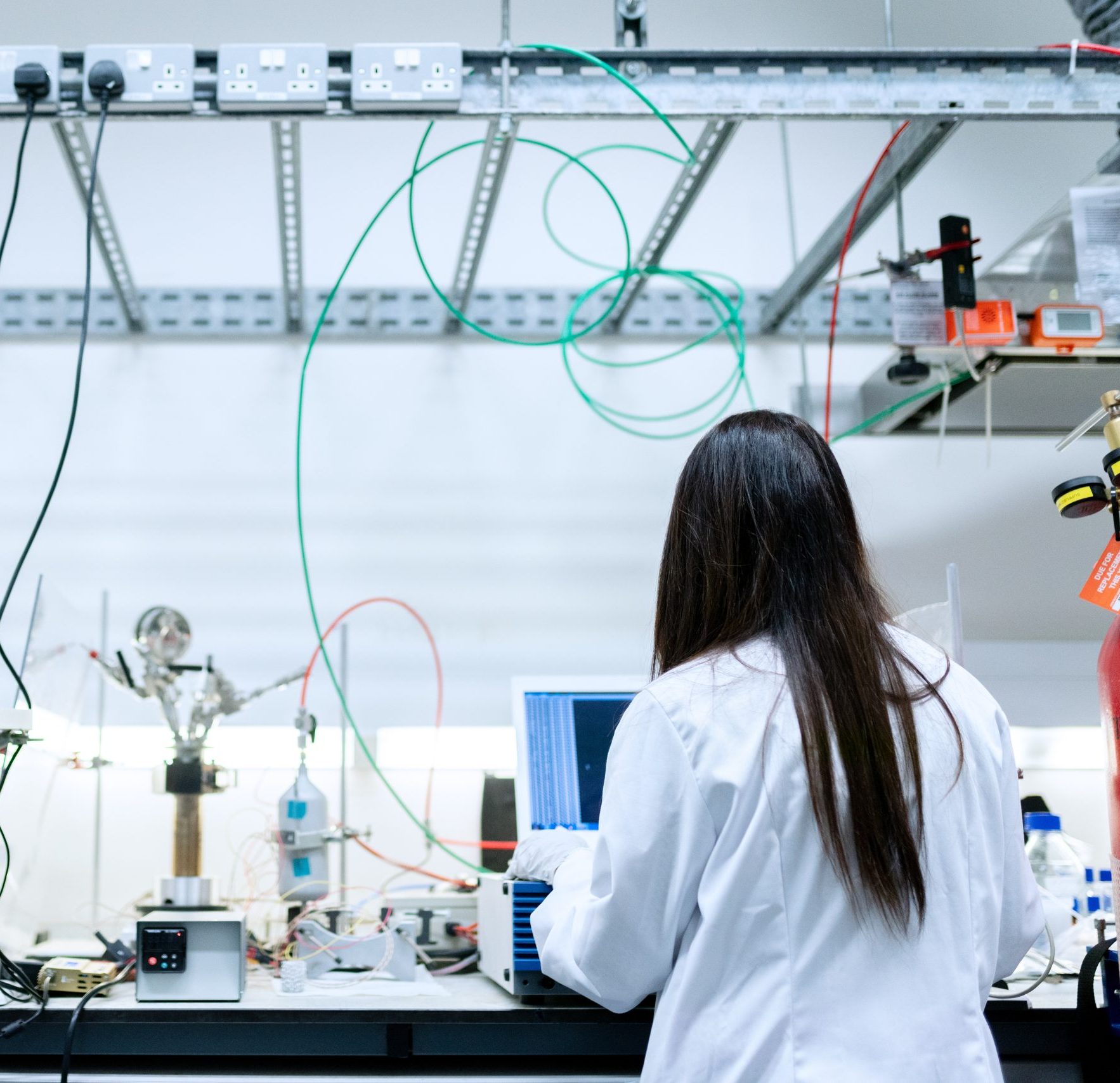
RNA Modification and Processing
Welcome to Transregio 319 RMaP
What is RMaP?
Transregio 319 RMaP is a joint venture combining:
Molecular Biology, Biochemistry, Cellular Biology, Chemical Biology, Biophysical Chemistry, Structural Biology, Developmental Biology, Genetics, Bioinformatics.
Structure of RMaP
A „Modificationslook for Processing“
B „Processing looks for Modifications“
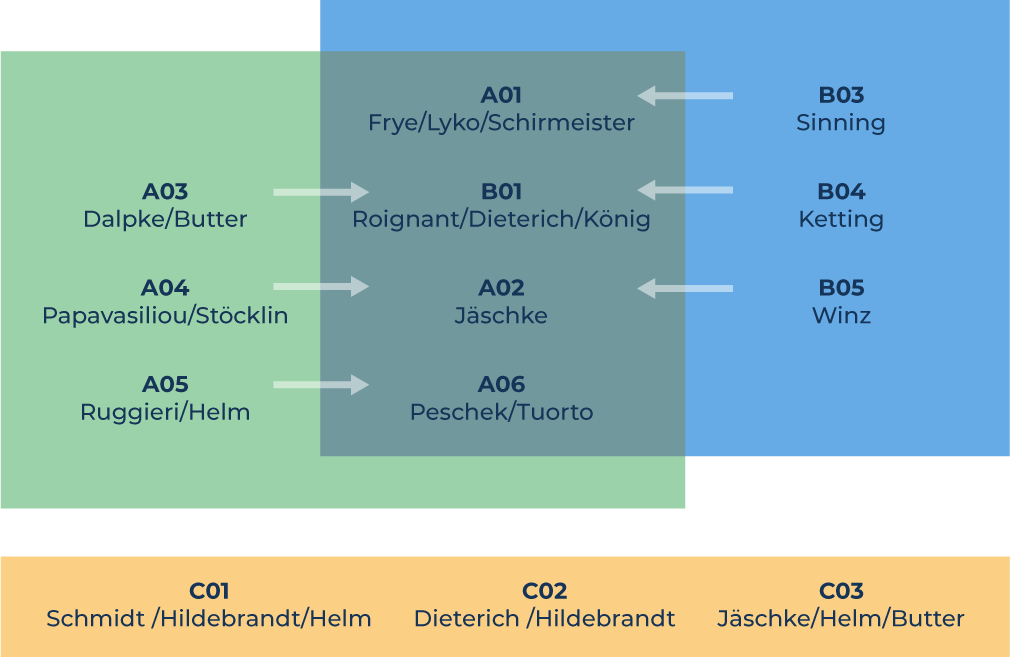
C „New RNA Technology for RMaP“
TRR Speaker Office

Prof. Dr. Mark Helm
Speaker

Dr. Marion Stemke
Executive officer and Coordinator

Claudia Alihodzic
Administrator TRR319

Doris Ebeling
Administration & Equal Opportunity officer

Doris Gebhardt
Administration & Equal opportunity officer
Members of the Scientific Board

Recent publications from the TRR 319
Podvalnaya N, Bronkhorst AW, Lichtenberger R, Hellmann S, Nischwitz E, Falk T, Karaulanov E, Butter F, Falk S, Ketting RF. piRNA processing by a trimeric Schlafen-domain nuclease. Nature. 2023 Oct;622(7982):402-409. doi: 10.1038/s41586-023-06588-2. Epub 2023 Sep 27. PMID: 37758951; PMCID: PMC10567574.
Motorin Y, Helm M. General Principles and Limitations for Detection of RNA Modifications by Sequencing. Acc Chem Res. 2023 Dec 8. doi: 10.1021/acs.accounts.3c00529. Epub ahead of print. PMID: 38065564.
Helm M, Bohnsack MT, Carell T, Dalpke A, Entian KD, Ehrenhofer-Murray A, Ficner R, Hammann C, Höbartner C, Jäschke A, Jeltsch A, Kaiser S, Klassen R, Leidel SA, Marx A, Mörl M, Meier JC, Meister G, Rentmeister A, Rodnina M, Roignant JY, Schaffrath R, Stadler P, Stafforst T. Experience with German Research Consortia in the Field of Chemical Biology of Native Nucleic Acid Modifications. ACS Chem Biol. 2023 Nov 14. doi: 10.1021/acschembio.3c00586. Epub ahead of print. PMID: 37962075.
Sung HM, Schott J, Boss P, Lehmann JA, Hardt MR, Lindner D, Messens J, Bogeski I, Ohler U, Stoecklin G. Stress-induced nuclear speckle reorganization is linked to activation of immediate early gene splicing. J Cell Biol. 2023 Dec 4;222(12):e202111151. doi: 10.1083/jcb.202111151. Epub 2023 Nov 13. PMID: 37956386; PMCID: PMC10641589.
Sun Y, Piechotta M, Naarmann-de Vries I, Dieterich C, Ehrenhofer-Murray AE. Detection of queuosine and queuosine precursors in tRNAs by direct RNA sequencing. Nucleic Acids Res. 2023 Nov 10;51(20):11197-11212. doi: 10.1093/nar/gkad826. PMID: 37811872; PMCID: PMC10639084.
Möhler M, Jäschke A. Future Perspectives for the Identification and Sequencing of Nicotinamide Adenine Dinucleotide-Capped RNAs. Acc Chem Res. 2023 Nov 7;56(21):3000-3009. doi: 10.1021/acs.accounts.3c00446. Epub 2023 Oct 18. PMID: 37852615; PMCID: PMC10634297.
Sun Y, Piechotta M, Naarmann-de Vries I, Dieterich C, Ehrenhofer-Murray AE. Detection of queuosine and queuosine precursors in tRNAs by direct RNA sequencing. Nucleic Acids Res. 2023 Oct 9:gkad826. doi: 10.1093/nar/gkad826. Epub ahead of print. PMID: 37811872.
Podvalnaya N, Bronkhorst AW, Lichtenberger R, Hellmann S, Nischwitz E, Falk T, Karaulanov E, Butter F, Falk S, Ketting RF. piRNA processing by a trimeric Schlafen-domain nuclease. Nature. 2023 Oct;622(7982):402-409. doi: 10.1038/s41586-023-06588-2. Epub 2023 Sep 27. PMID: 37758951; PMCID: PMC10567574.
A continuously updated publication list can be found here.






























